This past January I was able to attend rstudio::conf 2019 in Austin. I was there for the full week, taking the Building Tidy Tools workshop and attending as many of the main conference program sessions as possible.
Here is a collection of highlights/takeaways from the conference:
Shortcuts
Coming into rstudio::conf I had a few keyboard shortcuts memorized. Things like copying a line of code down (CMD + SHIFT + DOWN ARROW) … inserting the pipe operator (CMD + SHIFT + M) … and inserting a code chunk (CMD + OPTION + I).
But there are so many more … a few handy ones that I picked up:
- Maximize an Rstudio pane:
SHIFT + CTRL + 1(1,2,3,4 for each pane clockwise) - Build a package:
SHIFT + CMD + B - Load functions from package after building:
SHIFT + CMD + L
My friends (and fellow Charlottesville R Users) Jeff Boichuk and Steve Mortimer have also developed an R package (CrossR) that includes a data frame with all Rstudio shortcuts:
devtools::install_github("jeffboichuk/CrossR")
library(CrossR)
shortcuts## # A tibble: 122 x 4
## category description mac windows
## <chr> <chr> <chr> <chr>
## 1 Build Build All Shift + Cmd + B Ctrl + Shift + B
## 2 Build Check Package Shift + Cmd + E Ctrl + Shift + E
## 3 Build Compile PDF Shift + Cmd + K Ctrl + Shift + K
## 4 Build Devtools Load All Shift + Cmd + L Ctrl + Shift + L
## 5 Build Knit Document Shift + Cmd + K Ctrl + Shift + K
## 6 Build Preview HTML Shift + Cmd + K Ctrl + Shift + K
## 7 Build Roxygenize Package Shift + Cmd + D Ctrl + Shift + D
## 8 Build Test Package Shift + Cmd + T Ctrl + Shift + T
## 9 Console Console Clear Ctrl + L Ctrl + L
## 10 Debug Debug Breakpoint Shift + F9 Shift + F9
## # … with 112 more rowsfind_shortcuts("roxygen")## # A tibble: 2 x 3
## category description mac
## <chr> <chr> <chr>
## 1 Build Roxygenize Package Shift + Cmd + D
## 2 Source Editor Insert Roxygen Skeleton Shift + Alt + Cmd + RRStudio 1.2 features
The RStudio team is working to make the RStudio IDE an even more comprehensive data science toolkit. Version 1.2 introduces features to support multiple languages, including python, SQL and D3. Jonathan McPherson gave a tour of the new multilingual support:
https://resources.rstudio.com/rstudio-conf-2019/new-language-features-in-rstudio
I put together a trivial example of what this looks workflow looks like for passing R objects to python (and vice versa) from chunk-to-chunk in an R markdown document:
```{r setup}
library(reticulate)
use_python("/usr/bin/python", required = TRUE)
```
```{python}
import random
x = random.randint(1,99)
print x
```
```{r}
# access the object 'x' from python environment with py$x
py$x
y <- py$x^2
y
```
```{python}
# access the object 'y' from r environment with r.y
print r.y
import math
z = math.sqrt(r.y)
print z
```For a more complete example (including D3 and SQL chunks), take a look at the spenders.Rmd example:
https://github.com/jmcphers/rstudio-1.2-features/blob/master/spenders.Rmd
New language features in RStudio - Jonathan McPherson
Tidy eval
Programming with tidy eval can seem daunting. One of the sessions of the Building Tidy Tools workshop focused on tidy eval. That talk demystified the quoting/unquoting workflow. Maybe more importantly for me, I found out that Rstudio is compiling a list of use-cases/examples of tidy eval in the wild:
https://community.rstudio.com/t/interesting-tidy-eval-use-cases/21121
Working with names and expressions in your tidy eval code - Lionel Henry
separate_rows()
The tidyr separate_rows() function isn’t a new by any means, but it’s one that I picked up during a demo at the conference. This will be for handy “wide-to-long” transformations on data that is packed into a single column (and separated by a common delimiter):
library(tidyr)
dat <-
data.frame(
id = 1:5,
diagnosis = c(NA, "A,B,C", "D", "B,E,F", "A")
)
dat## id diagnosis
## 1 1 <NA>
## 2 2 A,B,C
## 3 3 D
## 4 4 B,E,F
## 5 5 Aseparate_rows(dat, diagnosis, sep = ",")## id diagnosis
## 1 1 <NA>
## 2 2 A
## 3 2 B
## 4 2 C
## 5 3 D
## 6 4 B
## 7 4 E
## 8 4 F
## 9 5 A.Rmd to .pptx
Again … not a new feature, but new to me:
https://bookdown.org/yihui/rmarkdown/powerpoint-presentation.html
You can convert R markdown to PowerPoint by specifying powerpoint_presentation as the output format in the YAML header:
---
title: "Figs"
author: "VP Nagraj"
date: "3/10/2019"
output: powerpoint_presentation
---nb this output format requires Pandoc 2.x
New packages
There were a ton of new (or at least new-ish) packages introduced at rstudio::conf.
ungeviz
https://github.com/wilkelab/ungeviz
Tools for visualizing uncertainty with ggplot2. This package is meant to provide helpful add-on functionality for ggplot2 to visualize uncertainty. The package is particularly focused on hypothetical outcome plots (HOPs) and provides bootstrapping and sampling functionality that integrates well with the ggplot2 API. The package name comes from the German word “Ungewissheit”, which means uncertainty.
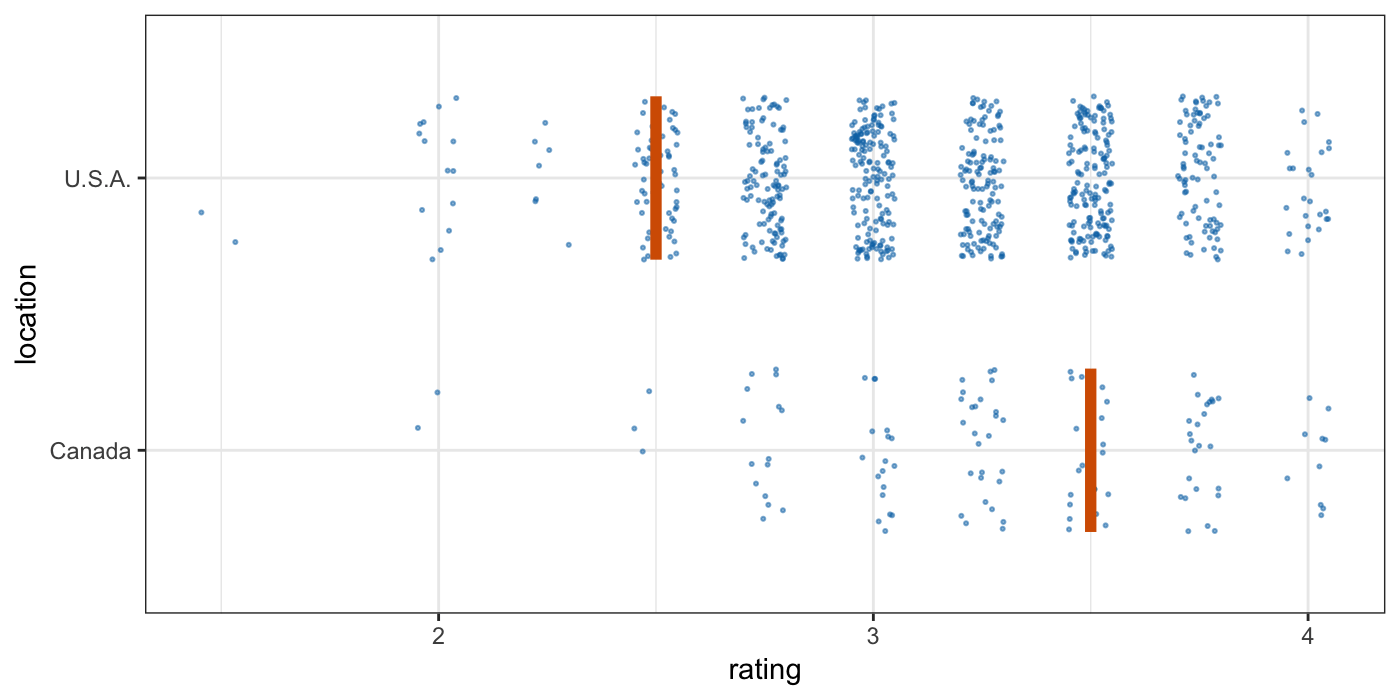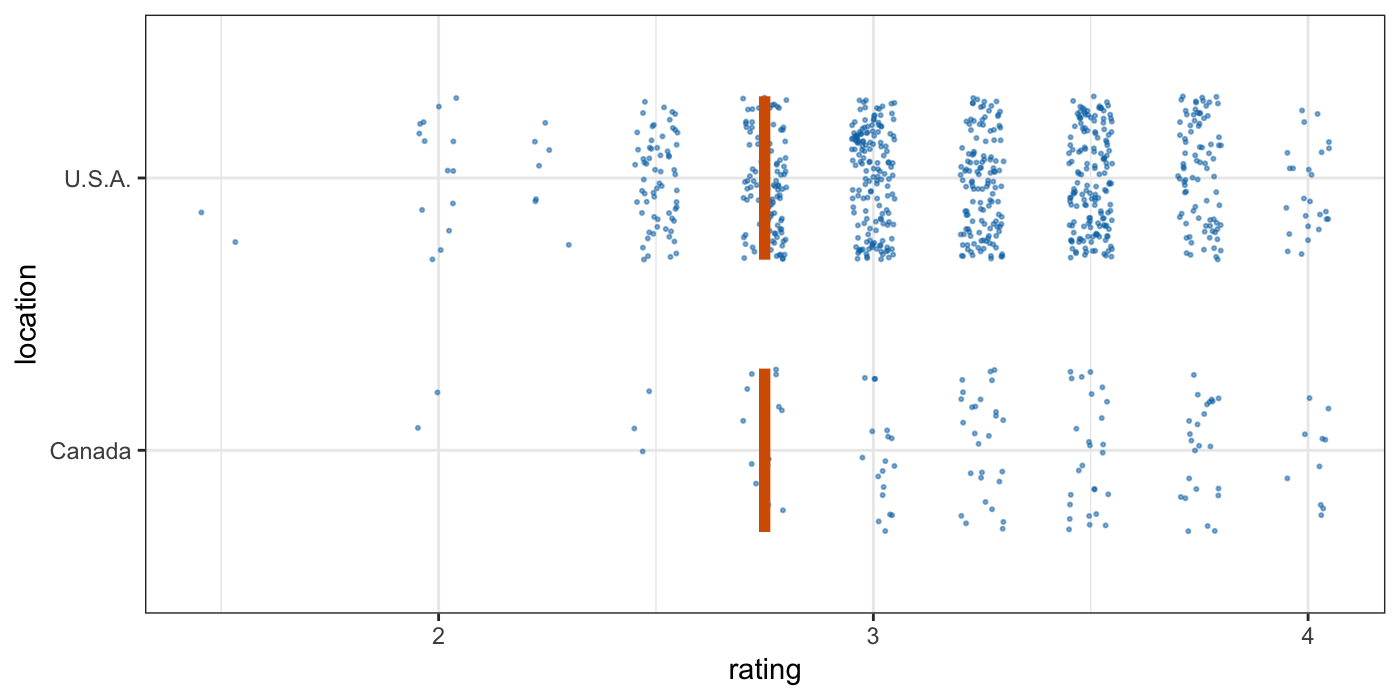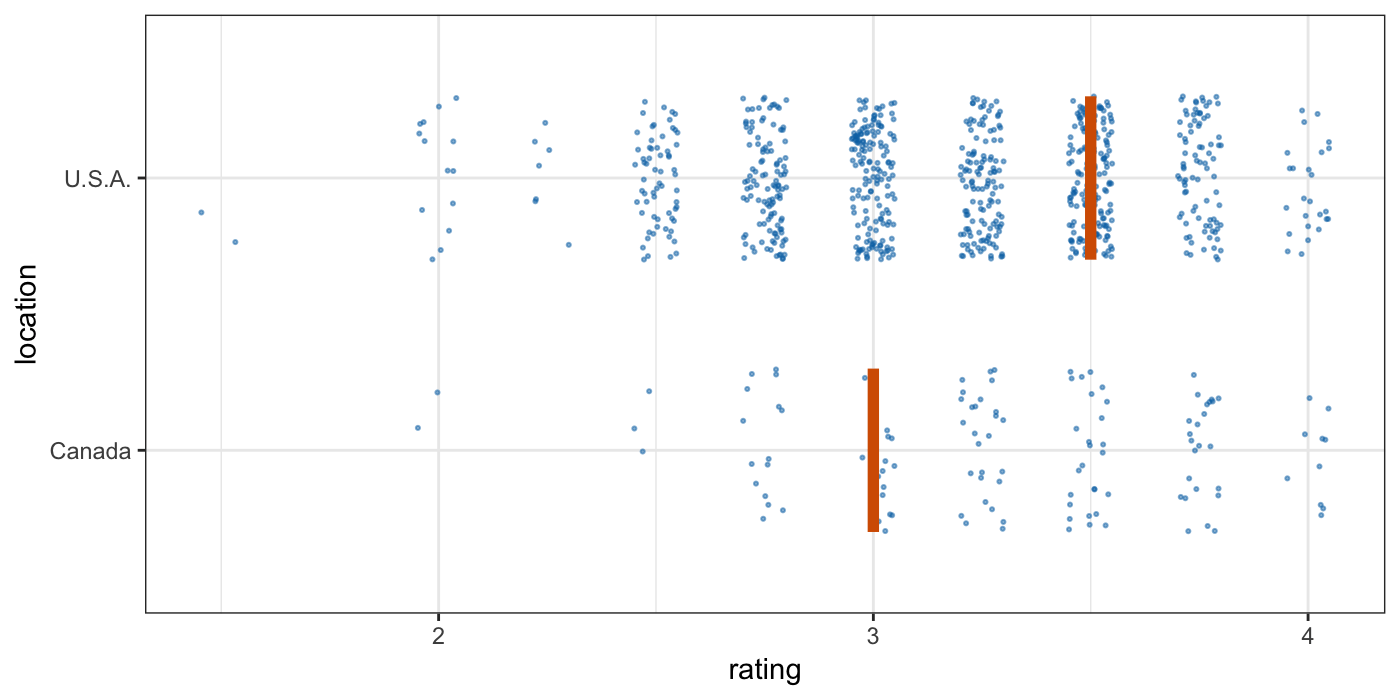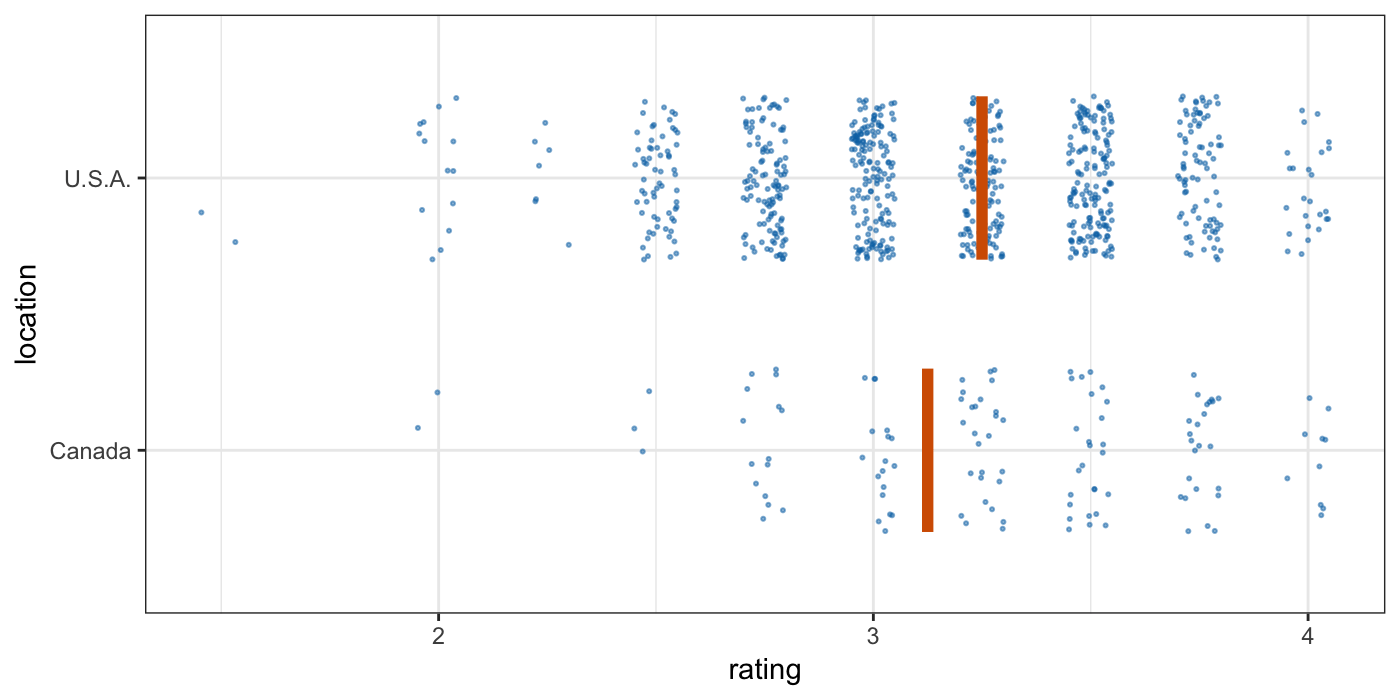
Visualizing uncertainty with hypothetical outcomes plots - Claus Wilke
tidymodels
https://github.com/tidymodels/tidymodels
tidymodels is a “meta-package” for modeling and statistical analysis that share the underlying design philosophy, grammar, and data structures of the tidyverse.
Solving the model representation problem with broom - Alex Hayes
gt
With the gt package, anyone can make wonderful-looking tables using the R programming language. The gt philosophy: we can construct a wide variety of useful tables with a cohesive set of table parts. These include the table header, the stub, the stub head, the column labels, the table body, and the table footer.
pagedown
https://github.com/rstudio/pagedown
Paginate the HTML Output of R Markdown with CSS for Print. You only need a modern web browser (e.g., Google Chrome) to generate PDF. No need to install LaTeX to get beautiful PDFs.

pagedown Creating beautiful PDFs with R Markdown and CSS - Yihui Xie






